FINAL PROJECTS

While completing all the background info for this website, I wanted to learn more about how the -13910 single base change affected people's expression of lactase (see Home-Genetic Causes for more information). It seemed likely that it was affecting the binding of a transcription factor and I determined that AP2 had a binding sequence that had a consensus sequence surrounding the -13910 bases. See left for my alignment of the AP2 consensus sequence followed by the lactose intolerant sequence and the lactose tolerant sequence around the -13910 location.
|
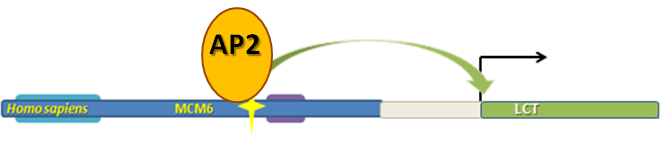
The image to the left is my interpretation of the AP2 transcription factor binding to the -13910 sequence in the MCM6 coding region/LCT promoter. Despite the image, which will be updated, the AP2 binding sequence is disrupted in those who are lactose tolerant indicating that AP2 would somehow inhibit lactase transcription.
|
METHODS
I would perform a ChIP assay plus qPCR to determine how much, if any AP2 is binding to the -13910 sequence [1]. To perform a ChIP assay, I have chosen to use an antibody that has been validated to work with ChIP, such as the AP2 antibody created by abcam. It would react with human cells and I would use cells from an intestinal biopsy of individuals with each combination of C and T (C/C, C/T, T/T) at the -13910 site. I would cross-link the protein to the DNA, probably with formaldehyde, and lyse the cells to extract the DNA and its binding factors. To see if AP2 is bound, I would digest the DNA enzymatically and perform an immunoprecipitation, also with an abcam kit to ensure proper functioning. Finally, the DNA must be eluted out, the cross-links need to be removed and the proteins need to be digested before the DNA can be sequenced. This sequence will be matched to the -13910 region of MCM6 and if it is a match, then we have a good indication that AP2 is binding to this region. A reason it may not bind would be that AP2 is generally seen as an activator and in this case, I am hypothesizing it would actually be repressing transcription.
If there is no AP2 detected, I propose to perform a DNA Pull Down Assay plus MudPIT mass spectrometry to determine what the the transcription factor is that is binding to this site. I would follow the protocol as defined by Robert A. Scott at the University of Georgia-Athens [2]. This protocol will help me determine which transcription factors are binding to this -13910 region of DNA, if any. It involves biotinylating the MCM6 DNA containing a wide-swath around the -13910 site. This will be incubated with a magnetic bead that binds biotin and also with cell extract (including transcription factors that may be binding). Since the bead is magnetic, it is possible to pull away the bead with attached DNA and bound proteins while washing away the rest of the cell extract. Finally, I would elute away the bound protein(s) and use Multi-Dimensional Protein Identification Technology (MudPIT) to analyze them. The University of Wisconsin-Madison Mass Spectrometry Facility has the capability to perform MudPIT which involves separating the proteins by molecular weight and ionization and then analyzing them in a mass spectrometer and comparing their results to known proteins.
I would perform a ChIP assay plus qPCR to determine how much, if any AP2 is binding to the -13910 sequence [1]. To perform a ChIP assay, I have chosen to use an antibody that has been validated to work with ChIP, such as the AP2 antibody created by abcam. It would react with human cells and I would use cells from an intestinal biopsy of individuals with each combination of C and T (C/C, C/T, T/T) at the -13910 site. I would cross-link the protein to the DNA, probably with formaldehyde, and lyse the cells to extract the DNA and its binding factors. To see if AP2 is bound, I would digest the DNA enzymatically and perform an immunoprecipitation, also with an abcam kit to ensure proper functioning. Finally, the DNA must be eluted out, the cross-links need to be removed and the proteins need to be digested before the DNA can be sequenced. This sequence will be matched to the -13910 region of MCM6 and if it is a match, then we have a good indication that AP2 is binding to this region. A reason it may not bind would be that AP2 is generally seen as an activator and in this case, I am hypothesizing it would actually be repressing transcription.
If there is no AP2 detected, I propose to perform a DNA Pull Down Assay plus MudPIT mass spectrometry to determine what the the transcription factor is that is binding to this site. I would follow the protocol as defined by Robert A. Scott at the University of Georgia-Athens [2]. This protocol will help me determine which transcription factors are binding to this -13910 region of DNA, if any. It involves biotinylating the MCM6 DNA containing a wide-swath around the -13910 site. This will be incubated with a magnetic bead that binds biotin and also with cell extract (including transcription factors that may be binding). Since the bead is magnetic, it is possible to pull away the bead with attached DNA and bound proteins while washing away the rest of the cell extract. Finally, I would elute away the bound protein(s) and use Multi-Dimensional Protein Identification Technology (MudPIT) to analyze them. The University of Wisconsin-Madison Mass Spectrometry Facility has the capability to perform MudPIT which involves separating the proteins by molecular weight and ionization and then analyzing them in a mass spectrometer and comparing their results to known proteins.
Below are the slides from the PowerPoint that I created for my Genetics 677 final project presentation, in a PDF file:
| final_presentation_file.pdf | |
| File Size: | 1493 kb |
| File Type: | |
REFERENCES
[1] "ChIP Assay Procedure and Essential Tools." Chromatin IP (ChIP Assays). Thermo Fischer Scientific, n.d. Web. 16 May 2013.
[2] Wang, Maiyao, Gina L. Lipscomb, Annette M. Keese, Gerrit J. Schut, Michael WW Adams, I. Jonathan Amster, Michael Thomm, and Robert A. Scott. N.d. MS. University of Georgia-Athens. Sept. 2007. Web. 16 May 2013. http://rscott.myweb.uga.edu/TR/STRES_final.pdf
[1] "ChIP Assay Procedure and Essential Tools." Chromatin IP (ChIP Assays). Thermo Fischer Scientific, n.d. Web. 16 May 2013.
[2] Wang, Maiyao, Gina L. Lipscomb, Annette M. Keese, Gerrit J. Schut, Michael WW Adams, I. Jonathan Amster, Michael Thomm, and Robert A. Scott. N.d. MS. University of Georgia-Athens. Sept. 2007. Web. 16 May 2013. http://rscott.myweb.uga.edu/TR/STRES_final.pdf
Page created by: Renae Geier, undergraduate student in genetics at the University of Wisconsin - Madison ([email protected])
Page last updated: 5/20/2013
Page last updated: 5/20/2013