PROTEIN (AMINO ACID) INFORMATION
Homo sapiens minichromosome maintenance complex component 6 (MCM6)
View Protein Sequence on the National Center for Biotechnology Information website
View Protein Sequence on the National Center for Biotechnology Information website
PROTEIN DOMAINS, FAMILIES, CONSERVED SITESMCM6 makes up one of six proteins that comprises the replicative helicase used in DNA replication (see Figure 1) [4]. The C-terminus end of the protein has a CBD, or Cdt1 binding domain, which is where the MCM6 (and thus whole helicase as well) interacts with the Cdt1 protein, which loads the helicase onto the DNA. Structural studies have shown that the CBD domain has a "winged-helix" (see Figure 1) [2]. I found this in a paper but not in any domain-finding websites, so I left it out of my diagram but I thought it was important to include nonetheless.
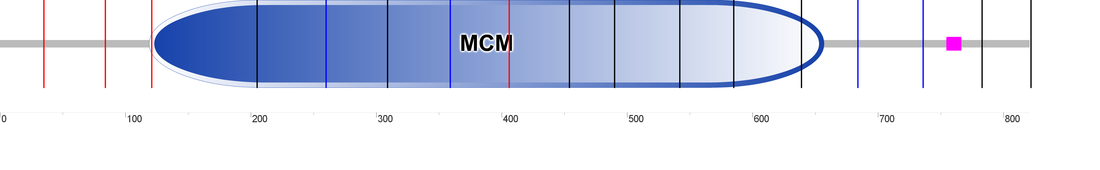
The mechanism of helicase loading and the details of how this protein complex interacts with DNA and the associated proteins has yet to be elucidated, but some of the suspected domains are listed below and shown in Figure 2. DOMAINS LISTED ON INTERPRO (see Figure 2 for their locations): Nucleic acid-binding, OB-fold (IPR012340) This domain binds to the single stranded nucleic acids (in DNA and RNA) or to oligosaccharides (a polymer of simple sugars). It is a five-stranded beta-barrel capped by an alpha-helix. Mini-chromosome maintenance, DNA-dependent ATPase (IPR001208) This domain is the catalytic center of the protein. This AAA+ protease domain hydrolyzes ATP which helps unwind double-stranded DNA during replication. Figure 3: These are the domains found by PFAM. The MCM_N domain correlates with the nucleic acid binding domain, and the MCM domain approximately correlates to the P-loop,as see in Figure 2. The MCM domain is found in other MCM proteins, numbers 2,3 and 5, all of which are a part of a hexamer (see Figure 1).
Figure 4: These are the domains found by SMART. The MCM domain is much wider here and includes both of the domains shown in Figures 2 and 3 (MCM_N/nucleic acid binding and MCM/P-loop).
|
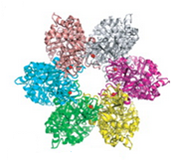
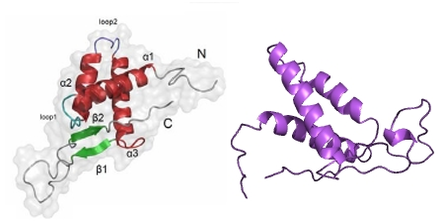
Figure 1: The hexamer structure of MCM 2-7 (left) [5]. The "winged-helix" structure of MCM6, determined by Wei et al., 2010 (right) [2].
Figure 2: MCM6 domains in humans and some homologs. For more information on these homologs, see my homolog page for more details.
FOR MORE INFORMATION ON THIS WEBSITE:
|
ANALYSIS AND DISCUSSION OF DOMAIN INFORMATION
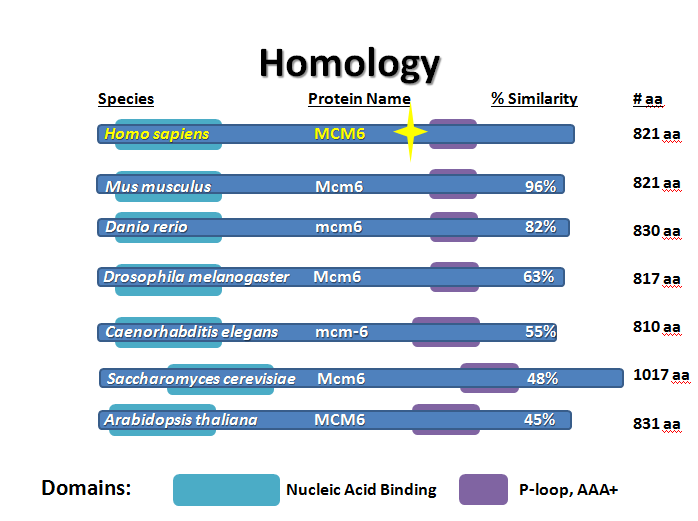
It makes sense how well conserved the MCM6 protein is in eukaryotes. Helicases are necessary to unwind and open the double-stranded DNA so that DNA polymerase can have access to the bases for replication. The domains are the same because for a helicase to function, it must be able to bind DNA and hydrolyze ATP so each MCM6 has these, as it is one of the helicase monomers. The differences in base conservation, although seemingly large (only 45% similar in Arabidopsis, see Homology page or Figure 2) is likely mostly silent and/or neutral substitutions or small differences for interacting with its specific proteins. I do not believe the function of MCM6 is significantly different across the homologs.
In addition, although bacterial, archaeic and eukaryotic helicases function similarly, their replication machinery and cell types are quite different so there is not an 'MCM6' homolog in the bacteria; their helicase is a dodecameric protein called DnaB but the archaeic helicase does have an MCM title [6]. I chose not to analyze in any detail the non-eukaryotic homologs because it was not significant to my project.
It makes sense how well conserved the MCM6 protein is in eukaryotes. Helicases are necessary to unwind and open the double-stranded DNA so that DNA polymerase can have access to the bases for replication. The domains are the same because for a helicase to function, it must be able to bind DNA and hydrolyze ATP so each MCM6 has these, as it is one of the helicase monomers. The differences in base conservation, although seemingly large (only 45% similar in Arabidopsis, see Homology page or Figure 2) is likely mostly silent and/or neutral substitutions or small differences for interacting with its specific proteins. I do not believe the function of MCM6 is significantly different across the homologs.
In addition, although bacterial, archaeic and eukaryotic helicases function similarly, their replication machinery and cell types are quite different so there is not an 'MCM6' homolog in the bacteria; their helicase is a dodecameric protein called DnaB but the archaeic helicase does have an MCM title [6]. I chose not to analyze in any detail the non-eukaryotic homologs because it was not significant to my project.
SEARCHES WHICH LED TO NO RESULTS
CHEMICAL GENETICS
In addition, there were no small molecules found that interact with MCM6, as portrayed here.
PARALOGS
I was also unable to find any paralogs, as portrayed here.
In addition, there were no small molecules found that interact with MCM6, as portrayed here.
PARALOGS
I was also unable to find any paralogs, as portrayed here.
REFERENCES
[1] "BioGPS Gene Report." BioGPS Blog. The Scripps Research Institute, 2013. Web. 06 Mar. 2013.
[2] Wei, Z., C. Liu, X. Wu, N. Xu, B. Zhou, C. Liang, and G. Zhu. "Characterization and Structure Determination of the Cdt1 Binding Domain of Human Minichromosome Maintenance (Mcm) 6." Journal of Biological Chemistry 285.17 (2010): 12469-2473. Web. 6 Mar. 2013.
[3] [1] "SMART: MCM Domain Annotation." SMART: MCM Domain Annotation. EMBL, 2012. Web. 21 Feb. 2013.
[4] Singleton, Martin R., Mark S. Dillingham, and Dale B. Wigley. "Structure and Mechanism of Helicases and Nucleic Acid Translocases." Annual Review of Biochemistry 76.1 (2007): 23-50. Annual Reviews. Web. 5 May 2013.
[5] Bochman, M. L., and A. Schwacha. "The Mcm Complex: Unwinding the Mechanism of a Replicative Helicase." Microbiology and Molecular Biology Reviews 73.4 (2009): 652-83. American Society for Microbiology. Web. 8 May 2013.
[6] Shin, J.-H. "The Replicative Helicases of Bacteria, Archaea, and Eukarya Can Unwind RNA-DNA Hybrid Substrates." Journal of Biological Chemistry 281.37 (2006): 26914-6921. The Journal of Biological Chemistry. The American Society for Biochemistry and Molecular Biology. Web. 16 May 2013.
[1] "BioGPS Gene Report." BioGPS Blog. The Scripps Research Institute, 2013. Web. 06 Mar. 2013.
[2] Wei, Z., C. Liu, X. Wu, N. Xu, B. Zhou, C. Liang, and G. Zhu. "Characterization and Structure Determination of the Cdt1 Binding Domain of Human Minichromosome Maintenance (Mcm) 6." Journal of Biological Chemistry 285.17 (2010): 12469-2473. Web. 6 Mar. 2013.
[3] [1] "SMART: MCM Domain Annotation." SMART: MCM Domain Annotation. EMBL, 2012. Web. 21 Feb. 2013.
[4] Singleton, Martin R., Mark S. Dillingham, and Dale B. Wigley. "Structure and Mechanism of Helicases and Nucleic Acid Translocases." Annual Review of Biochemistry 76.1 (2007): 23-50. Annual Reviews. Web. 5 May 2013.
[5] Bochman, M. L., and A. Schwacha. "The Mcm Complex: Unwinding the Mechanism of a Replicative Helicase." Microbiology and Molecular Biology Reviews 73.4 (2009): 652-83. American Society for Microbiology. Web. 8 May 2013.
[6] Shin, J.-H. "The Replicative Helicases of Bacteria, Archaea, and Eukarya Can Unwind RNA-DNA Hybrid Substrates." Journal of Biological Chemistry 281.37 (2006): 26914-6921. The Journal of Biological Chemistry. The American Society for Biochemistry and Molecular Biology. Web. 16 May 2013.
Page created by: Renae Geier, undergraduate student in genetics at the University of Wisconsin - Madison ([email protected])
Page last updated: 8/26/2013
Page last updated: 8/26/2013